(openPR) In a technological tour de force, Berlin scientists have created a virtual model of an early fly embryo. Its interactive interface allows researchers to explore the blueprint that underlies development at unprecedented spatial resolution and predict which cells express which genes.
---
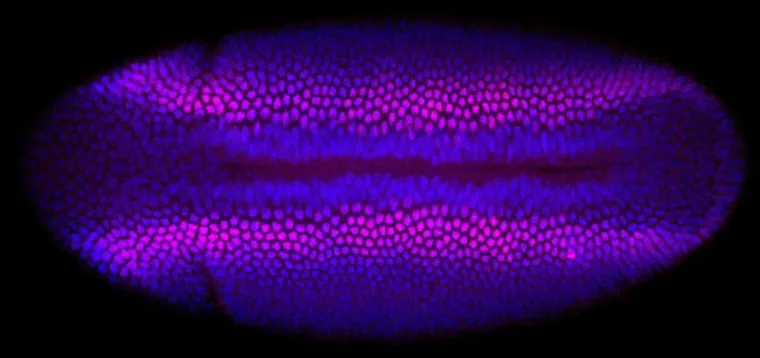
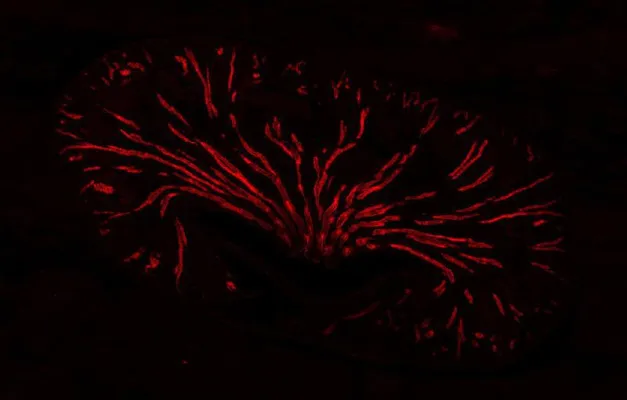
After 13 rapid divisions a fertilized fly egg consists of about 6,000 cells. They all look alike under the microscope. However, each cell of a Drosophila melanogaster embryo already knows by then whether it is destined to become a neuron or a muscle cell – or part of the gut, the head, or the tail. Now, Nikolaus Rajewsky’s and Robert Zinzen’s teams at the Berlin Institute of Medical Systems Biology (BIMSB) of the Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) have analyzed the unique gene expression profiles of thousands of single cells and reassembled the embryo from these data using a new spatial mapping algorithm. The result is a virtual fly embryo showing exactly which genes are active where at this point in time. “It is basically a transcriptomic blueprint of early development,” says Robert Zinzen, head of the Systems Biology of Neural Tissue Differentiation Lab. Their paper appears as a First Release in the online issue of Science.
“Only recently has it become possible to analyze genome-wide gene expression of individual cells at a large scale. Nikolaus recognized the potential of this technology very early on and established it in his lab,” says Zinzen. “He started to wonder whether – given a complex organized tissue – one would be able to compute genome-wide spatial gene expression patterns from single-cell transcriptome data alone.” BIMSB combines laboratories with different backgrounds and expertise, emphasizing the need of bringing computing power to biological problems. It turns out the institute had not only the perfect model system – the Drosophila embryo – to address Rajewsky’s question, but also the right people with the right expertise, from physics and mathematics to biochemistry and developmental biology.
“The virtual embryo is much more than merely a cell mapping exercise,” says Nikolaus Rajewsky, head of the Systems Biology of Gene Regulatory Elements Lab, who enjoyed returning to fly development 15 years after studying gene regulatory elements in Drosophila embryos during his post-doctoral time at the Rockefeller University. Using the interactive Drosophila Virtual Expression eXplorer (DVEX) database (www.dvex.org), researchers can now look at any of about 8,000 expressed genes in each cell and ask, “Gene X, where are you expressed and at what level? What other genes are active at the same time and in the same cells?” It also works with the enigmatic long non-coding RNAs. “Instead of time-consuming imaging experiments, scientists can do virtual ones to identify new regulatory players and even get ideas for biological mechanisms,” says Rajewsky. “What would normally take years using standard approaches can now be done in a couple of hours.”
Breaking the synchronicity of the first cell divisions
In their paper, the MDC researchers describe a dozen new transcription factors and many more long non-coding RNAs that have never been studied before. Also, they propose an answer to a question that has puzzled scientists for 35 years: How does the embryo break synchronicity of cell divisions to develop more complex structures?
In a process called gastrulation, distinct germ layers form and cells become restricted with regard to which tissues and organs they may differentiate into. “We believe that the Hippo signaling pathway is at least partly responsible for setting up gastrulation,” says Rajewsky. The pathway controls organ size, cell cycles and cell proliferation, but had never been implicated in the development of the early embryo. “We not only showed that Hippo is active in the fly, but we could even predict in which regions of the embryo this would lead to a different onset of mitosis and therefore break synchronicity. And that is just one example for how useful our tool is to understand mechanisms that have escaped traditional science.”
Project underwent a tough gestation period
When the researchers started creating the virtual embryo, they did not know whether it would be possible. A key pillar of their eventual success is the Drop-Seq technology, a droplet-based, microfluidic method that allows the transcriptional profiling of thousands of individual cells at low cost. This technique had been newly set up in the Rajewsky lab by Jonathan Alles, a summer student.
However, the fly embryos needed to be selected precisely at the onset of gastrulation. Philipp Wahle, a PhD student in Robert Zinzen’s lab, hand-picked about 5,000 of them before dissociating them into single cells. “I was convinced this would give us a large and completely unique data set. This was a great motivation for me,” says Wahle. That laborious process created a new challenge. “You need to collect over several sessions to have enough material for a sequencing run,” says Christine Kocks, who led the single-cell sequencing team. It was composed of Jonathan Alles, Salah Ayoub and Anastasiya Boltengagen, who jointly with computational scientist
Nikos Karaiskos optimized the droplet-based sequencing. “So we had to find a way to stabilize the transcriptomes in the cells,” added Kocks. “Finally, based on his earlier work with C. elegans embryos, Nikolaus suggested using methanol.” The new single-cell fixation method was published in BMC Biology in May 2017.
As the data got better and better, Nikos Karaiskos, a theoretical physicist and computational expert in Rajewsky’s lab, took on the challenge of spatially mapping such a large number of cells to their precise embryonic position. None of the existing approaches in the field of spatial transcriptomics was suitable to reconstruct the Drosophila embryo. “It was a reiterative process to filter the data, see what is inside and try to map it. It changed many times along the way,” says Karaiskos. There was a lot of back and forth between members of the computer lab and wet lab – exchanges that are a defining characteristic of the BIMSB. “I had to question my work all the time, see where it was lacking and develop something better.” He came up with a new algorithm called DistMap that can map transcriptomic data of cells back to their original position in the virtual embryo.
Navigating unchartered territory
The construction of the virtual embryo allowed Karaiskos to readily predict the expression of thousands of genes, an almost impossible task by traditional experimental means. Philipp Wahle, supported by Claudia Kipar, validated these predictions by visualizing the gene expression profiles at the bench with a traditional approach: In situ hybridization allows visualizing patterns of gene expression with colorful dyes that are visible under the microscope. “At this stage, a single layer of cells surrounds the entire fly embryo,” says Wahle. “This makes it very accessible, thus enabling you to compare the computational data with imaging.”
It is the first time that it has been possible to look at the about 6,000 cells of the embryo individually, assess their gene expression profiles – and understand what determines their behavior in the embryo. “The most important technological advance of this study is that we don’t lose the spatial information that is required to understand how embryonic cells act in concert,” say the scientists. “This really is unchartered territory and requires new bioinformatics approaches to make sense of the collected data. This worked beautifully in our collaboration, not least because of the unique make-up of the Rajewsky lab, which integrates wet lab and computational approaches.” One major advantage is that both groups are not only interested in technology but have specific biological questions that motivate them, says Rajewsky. “Robert has a deep understanding of early development. We can do single-cell sequencing runs and have the computational power to develop the tools that help us actually understand the underlying gene regulatory interactions.”
The groups are already planning follow-up projects. One example would be to map the cells at different time points to see how they work together to form organs and tissues. Another would be to check whether the mapping approaches are applicable to more complex tissues.
Karaiskos, N. et al. (2017): “The Drosophila embryo at single-cell transcriptome resolution.” Science. doi:10.1126/science.aan3235
Weitere Informationen:
- http://dx.doi.org/10.1126/science.aan3235 – The publication at the Science website
- https://insights.mdc-berlin.de/en/2017/08/reconstructing-life-at-its-beginning-cell-by-cell/ Online version of this press release
- http://www.dvex.org/ Drosophila Virtual Expression eXplorer
- https://www.mdc-berlin.de/1151037 Rajewsky Lab’s website
- https://www.mdc-berlin.de/40467197 Zinzen Lab’s website
Quelle: idw